Dilfuza Djamalova
Projects
Ongoing projects
AMR prediction in clinical isolates | July 2024 - Present
| ML-based trimming of nanopore sequences | July 2024 - Present |
PhD projects
Feature extraction pipeline | March 2021 - Present
- Built a snakemake pipeline to extract sequence-based features from bacterial genomes
- Pipeline takes the genome data of the bacterial strains as input and produces a comprehensive table with the computed features for each gene as an output
- Pipeline implements egnogg-mapper, esearch, FreeSASA, Foldseek, CodonW, and GOATOOLS along with custom python scripts
- Pipeline can be run with command
snakemake -p combine_features -j --cores 36 --use-conda, where-pspecifies the rule name,-jallows parallel execution of the non-connected rules,--coresallocates the required number of CPUs,--use-condaallows the python scripts or the required software to be executed in its conda environment.
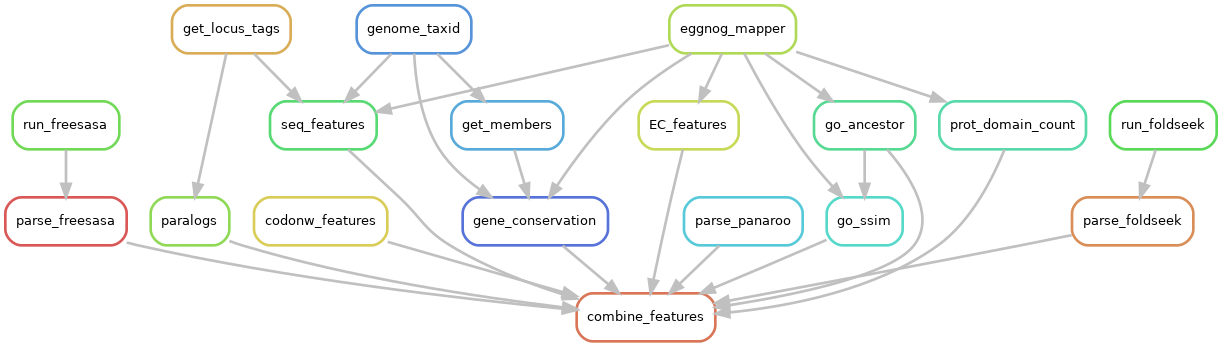 Fig. Rulegraph of the feature extraction pipeline.
Fig. Rulegraph of the feature extraction pipeline.
Bacterial fitness prediction | March 2021 - Present
- Implemented classifiers (XGBoost, Logistic Regression, Random Forest) using the Hydra Python framework.
- Pipeline architecture facilitates model training by allowing configuration files and data pre-processing steps (e.g. imputation of missing values) to be switched from the command line. It also allows new data processing or model training steps to be added without rewriting a significant amount of code.
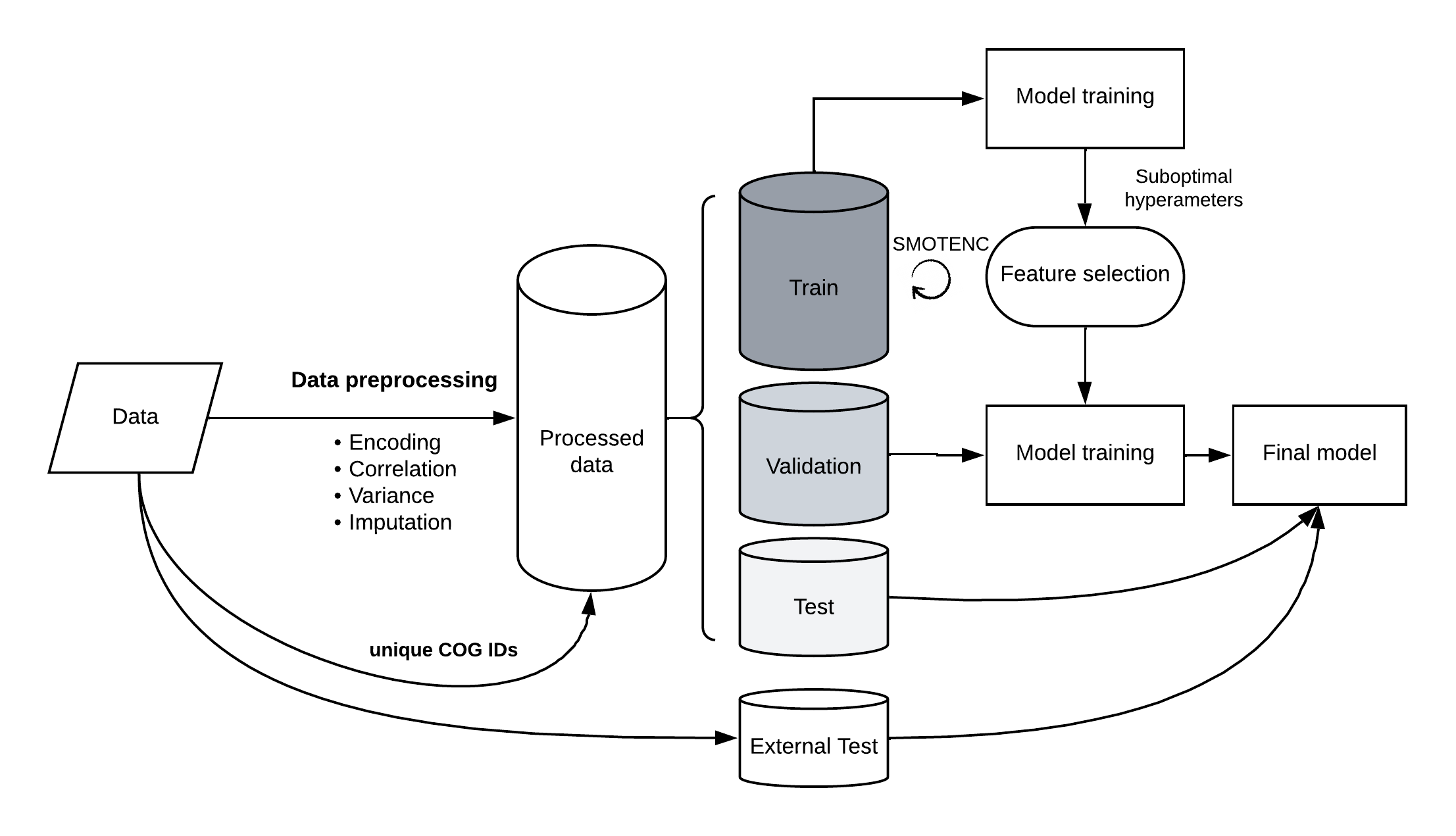 Fig. Model training simplified scheme.
Fig. Model training simplified scheme.
Code snippets used to generate plots | March 2021 - Present
- This repo contains Jupyter notebooks and Python scripts that are themselves small projects, e.g. to perform an incremental strain inclusion analysis, to assess a predictive power of sequence-based features, and to perform a cross-taxonomy prediction of bacterial essential genes at species, genera, family, and phylum levels.
- Also, Jupyter notebooks (1, 2, 3) contains example scripts to generate various plots.
Master’s project
Code snippets used in Master’s project | Sep 2018 - June 2020
- This repo contains python code snippets used in my Master’s project titled Re-classification of species and genera in family of Bacillaceae.
- Code snippets can be used to mainly collect bacterial genomes from NCBI Refseq/Genbank.
Coding practices
- I regularly try to update this repo with the solutions to ROSALIND tasks
- Repo contains Jupyter notebooks named after ROSALIND sections, e.g.
Python VillageorBioinformatics Stronghold.